
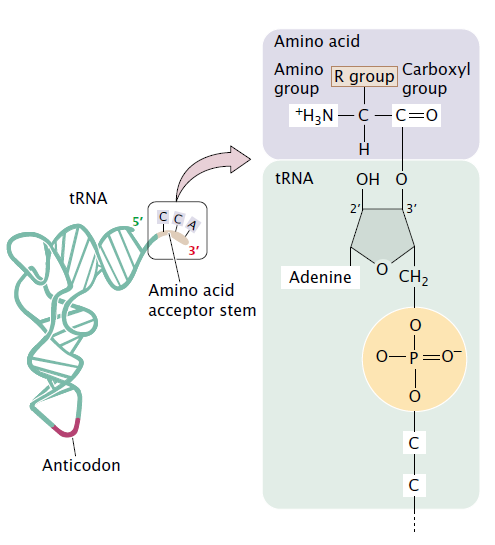
It has no way of checking each tRNA is matched with. When a ribosome pairs a 'CGC' tRNA with 'GCG' codon, it expects to find an alanine carried by the tRNA. Aspartyl-tRNA synthetase (blue and green) bound to tRNA (red).
Available from: ī Nagaoka, Yoshiyuki, et al. Aminoacyl-tRNA synthetases ensure that the proper amino acids are used to build proteins. Section 29.2, Aminoacyl-Transfer RNA Synthetases Read the Genetic Code. The threonine-AMP is shown in green.Ī Berg JM, Tymoczko JL, Stryer L.
The editing site can recognize serine and cleave it from the tRNA.įigure: Interaction network of the activation site of the threonyl-tRNA synthetase in A. For this purpose, aminoacyl-tRNA synthetases have a second, editing site that proofreads the charged tRNA. Nonetheless, the threonyl-tRNA synthetase could still be mischarged with serine because the zinc ion site is not specific enough, leading to 1% mischarged tRNAs. Uniquely to threonyl-tRNA synthetase, it contains a Zn 2+ ion that exhibits an interaction structure specific to threonine. Ribosomes are required for the translation of the genetic information in mRNA into a polypeptide chain. Fistly, the amino acid must be activated which involves the addit. We review their content and use your feedback to keep the quality high. This bond provides the energy for the synthesis of the peptide bond. How does an amino acid become attached to a tRNA what mechanism ensures amino acids get attached to the correct tRNA Who are the experts Experts are tested by Chegg as specialists in their subject area. The amino acid is attached at the 3 end of tRNA with a high energy bond forming charged tRNA. Valine has a methyl group instead of the hydroxy group of threonine, and serine, on the other side, has no additional methyl side group. The enzyme has a three part active site that recognizes three smaller molecules, a specific amino acid, ATP and a specific tRNA. Hydrophobic amino acids such as glutamine and proline are not found on the surface. For example, threonine, catalyzed by threonyl-tRNA synthetase, is very similar to valine and serine. mRNA AUG CAC UCU GCC GAU AAC CCC UGG UUU GAG UUC GGG AGA. First, the activation site, where the amino acid binds, constitutes a complex network of intermolecular interactions. The rules governing the tRNA-amino acid specificity depend on the codon on the tRNA, and the correct incorporation of the amino acid by the aminoacyl-tRNA synthetase enzyme. Origins of Life and Evolution of Biospheres Springer Journals Aminoacyl-tRNA sythetases are highly specific to their corresponding amino acid. Alone, an amino acid is not the substrate necessary to allow for the formation of peptide bonds. The nine hydrophilic amino acids are listed below, with the remaining two. The nine hydrophobic amino acids are alanine (Ala), glycine (Gly), valine (Val), leucine (Leu), isoleucine (Ile), phenylalanine (Phe), proline (Pro), methionine (Met), and tryptophan (Trp). The aa-tRNA, along with particular elongation factors, deliver the amino acid to the ribosome for incorporation into the polypeptide chain that is being produced during translation. Of the 20 common amino acids, all are defined by their R group's chain atoms.
#How do hydrophobic amino acids get to a trna code#
We interpret these results considering modelsfor the origin and evolution of the genetic code in which an initial version, containing fewer amino acids, was modified by the incorporationof new amino acids following duplication and divergence of previoussynthetases and tRNA molecules. Aminoacyl-tRNA (also aa-tRNA or charged tRNA) is tRNA to which its cognate amino acid is chemically bonded (charged). We argue that the evolution of aminoacyl-tRNA synthetases was determined by the characteristics of their corresponding amino acids. Furthermore, the phylogenetic trees of Classes I and II enzymes are highly correlated with dendrograms obtained for their cognate amino acids by using the indices in the AAIndex database.
The division of the enzymes in Classes I and II follows to a great extent a division in the chemical and biological properties of their cognate amino acids. The division of the aminoacyl-tRNA synthetases in two classes is compared with a division of the amino acids in two classes, obtained from the AAIndex databank by a principal component analysis. On the Classes of Aminoacyl-tRNA Synthetases, Amino Acids and the Genetic Code On the Classes of Aminoacyl-tRNA Synthetases, Amino Acids and the Genetic CodeĬavalcanti, Andre Soares Leite, Elisa Neto, Benício Ferreira, Ricardo
